In an earlier post I showed how plotSimmap(...,type="fan") could be used to plot densityMap or contMap style plots using the object of class "densityMap" or "contMap" returned invisibly by each function, respectively.
Well, I have now build this directly into contMap & densityMap (and the S3 generic plot for objects of class "densityMap" and "contMap"). Check it out:
> require(phytools)
Loading required package: phytools
> packageVersion("phytools")
[1] ‘0.2.76’
> contMap(tree,x,type="fan",fsize=0.9)
Loading required package: phytools
> packageVersion("phytools")
[1] ‘0.2.76’
> contMap(tree,x,type="fan",fsize=0.9)
> X<-densityMap(mtrees,outline=TRUE,fsize=0.9)
sorry - this might take a while; please be patient
sorry - this might take a while; please be patient
Argh - way too cluttered. Let's try a circular tree instead!
> class(X)
[1] "densityMap"
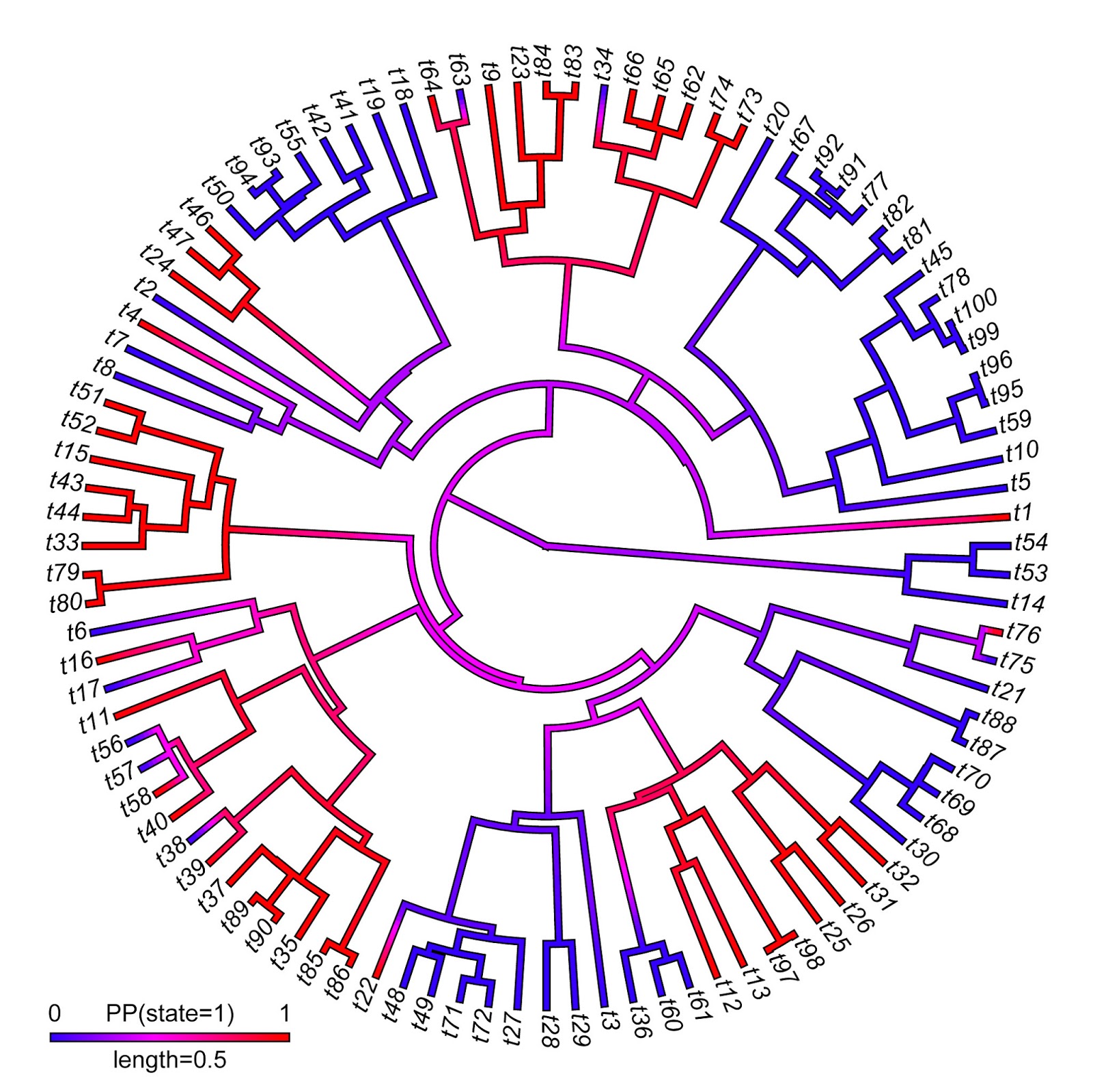
> plot(X,type="fan",fsize=0.9,outline=TRUE)
[1] "densityMap"
> plot(X,type="fan",fsize=0.9,outline=TRUE)
Since these updates involved changes to a number of methods in phytools - the best bet is to update to the latest non-CRAN phytools build (phytools 0.2-76).
That's it.


Hello Liam, greetings from Costa Rica
ReplyDeleteI've been trying to run a fan tree, circular tree, with features, but as a result I get the circular tree but without the species label and I get the following error:
Error in rep(" ", offset) : invalid 'times' argument
I copy the tree code
tree<- tree3
svl<- setNames(dat[,1], row.names(dat))
obj<-setMap(contMap(tree,svl, plot = F), invert=T)
plot(obj,fsize=c(0.4,0.8),outline=FALSE,lwd=c(1,4),leg.txt="Digestibility (mod mix)", type="fan")
Do you know what the problem might be? I have the latest version of R and updated all the packages
Thank you very much.
This was a bug in the previous version of phytools, but has already been fixed on CRAN.
DeleteHi Liam,
ReplyDeleteThank you so much for the very useful phytools package.
I'm also having the same problem as Jorge. I can do this plot
plotTree.wBars(obj$tree,count_andro,method="plotTree",
tip.labels=FALSE,fsize=1.5,col=cols,scale=0.004, border='lightgrey',width = 1)
However, when I specified that I want a fan tree, it doesn't work
plotTree.wBars(obj$tree,count_andro,method="plotTree",
tip.labels=FALSE,fsize=1.5,col=cols,scale=0.004, border='lightgrey',width = 1, type="fan")
and throws an error:
Error in rep(" ", offset) : invalid 'times' argument
I have looked for this error in the internet, but I haven't found useful information for this specific case.
Thank you so much in advance!
Wendy
Wendy. This bug is from a previous phytools version & should already be fixed on CRAN.
Delete@Valenatura and Jorge, you can solve this problem by adding the argument 'offset=1' to your plot() or contMap() call. @Liam, I guess this is a bug in plotFan, but I cannot figure out why it doesn't work when there is the if(is.null(offset)) offset <- 1 line in there that should overcome it.
ReplyDeleteHave you tried updating phytools from CRAN? I believe this bug has been fixed.
Delete