A recent R-sig-phylo subscriber posted two questions (1, 2) about connecting the tips of a plotted tree to mapped lat/long coordinates on a global map. He was certainly on the right track, but I just posted a better solution (code for the full function, here). I don't have too much time to get into the details on how this was done, but it uses code from my simple tree plotter (with x & y flipped, of course); and plot the map & tree in the same plotting object.
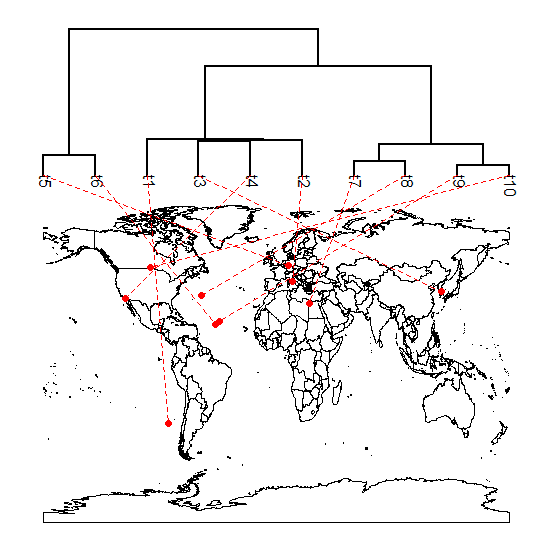
Here is an example of a result from a simulated tree & the submitted lat & long coordinates of the R-sig-phylo reader:
Because this was the user request, and I don't have much time to work on this kind of stuff this week - I have only created this plotter to use a full, global map; however, it would be pretty straightforward (I think) to generalize it to an arbitrary map by pulling the map coordinates from the map object in memory.
No comments:
Post a Comment
Note: due to the very large amount of spam, all comments are now automatically submitted for moderation.