I just developed a new "static" 3D phylomorphospace function; i.e., one that uses R base graphics to simulate three dimensions in a static plot, rather than the rgl library. The advantage of a static plot over a (much more impressive looking) spinning dynamic plot is that it is much easier to modify the plot using base graphics, and it can be exported in standard formats or combined with other plots. Code for this new version is here. The method uses scatterplot3d internally. It is also in a new phytools build (phytools 0.3-40) which can be installed from source.
Here's a quick demo of each method and the different products they create:
Loading required package: phytools
> packageVersion("phytools")
[1] ‘0.3.40’
> # simulate tree & data
> tree<-pbtree(n=26)
> tree$tip.label<-LETTERS
> X<-fastBM(tree,nsim=3)
> # first, method="dynamic" (the default)
> xx<-phylomorphospace3d(tree,X)
> movie3d(xx,duration=20,dir=".",movie= "phylomorphospace3d")
Will create: ./phylomorphospace3d.gif
Deleting frames.
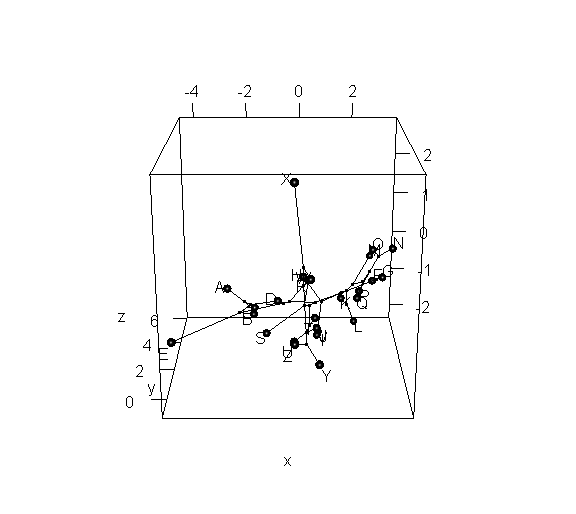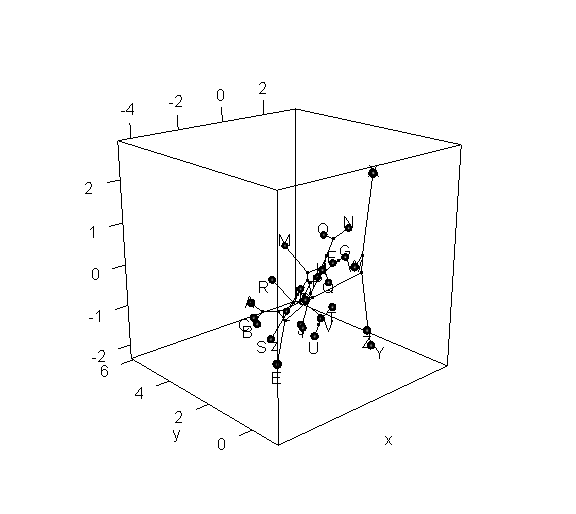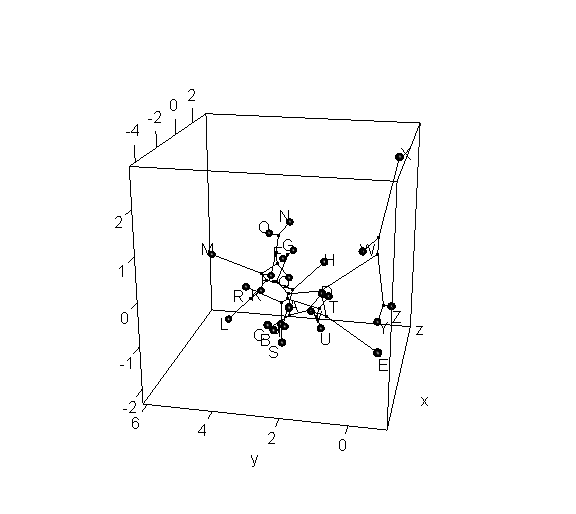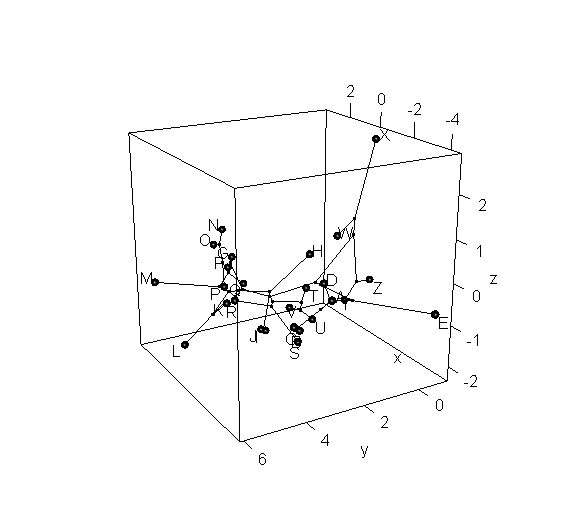
> phylomorphospace3d(tree,X,method="static")
> phylomorphospace3d(tree,X,method="static",angle=-30)
Obviously, the dynamic plot makes it a lot easier to see depth in the 3D plot; but the flat plot has the advantage of being something that we can more easily print to a page or combine with other R graphics.
That's it.

Dear Liam,
ReplyDeleteI tried to install the newest version phytools because I need to make 3D static graph (I will put it in my next scientific paper), but I did not have any luck.
Error message was:
"Error in read.dcf(file.path(pkgname, "DESCRIPTION"), c("Package", "Type")) :
cannot open the connection
In addition: Warning messages:
1: In unzip(zipname, exdir = dest) : error 1 in extracting from zip file
2: In read.dcf(file.path(pkgname, "DESCRIPTION"), c("Package", "Type")) :
cannot open compressed file 'phytools_0.3-40.tar.gz/DESCRIPTION', probable reason 'No such file or directory'"
I tried to do it manually (copied folder to R library but that did not help).
Do you have any idea what I am doing wrong?
Also I am trying to turn of taxa labels but without luck...
Thanks in advance!
Tanja Vukov
Hi Tanja.
DeleteI believe this method is in the latest CRAN version (phytools 0.3-72), so you should just be able to install directly from CRAN, i.e.:
install.packages("phytools")
and then select a mirror repository.
Let me know if that does not work.
All the best, Liam
This comment has been removed by the author.
ReplyDeleteDear Liam,
ReplyDeleteI know this post is a few years old, but I was wondering if you can assist me with some issues I'm having with using fancyTree() to create a static 3D phylomorphospace plot. The resulting plot is very "squished" and I can't seem to make the margins smaller so that the plot itself is visible. I've tried adding a mar= argument like in scatterplot3d() but it doesn't change anything. I've also tried to reset my graphics default margins, but that does not seem to help either.
Additionally, I was wondering if it is possible to add phylogenies to an existing plot in order to show multiple in the same plot?
Thank you so much,
Cecina
Your site is very informative and your articles are wonderful.www.123moviestube.io
ReplyDeleteThis comment has been removed by the author.
ReplyDelete