A user recently submitted the following question:
is it possible to create a phenogram (that is, a projection of the tree into a two dimensional space defined by morphology on the vertical axis and time, see here) with different clades as different colors?
The function phenogram already allows a mapped discrete character, so if we want to color arbitrary clades on our tree different colors, we just need to combine phenogram with the phytools function paintSubTree, described here.
Let's try it in the following illustrating example:
> # first simulate a random tree
> tree<-pbtree(n=20,scale=100)
> # now plot the tree with node labels
> # so we can select the clades we want to paint
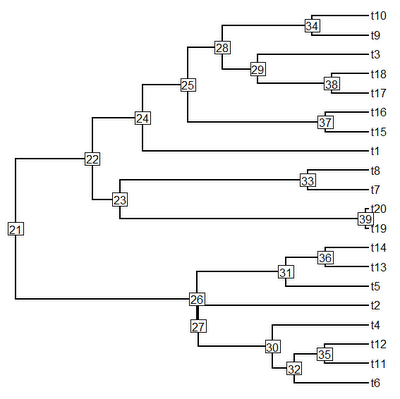
> plotTree(tree,pts=F,node.numbers=T)
OK, this tree has two main clades: one descended from node number "22", and the other descended from node number "26". Let's first paint the former clade with state "2" and the latter clade with state "3", we can leave the stem branches of each clade in state "1" (although we need not).
> tree<-paintSubTree(tree,node=22,state="2")
> tree<-paintSubTree(tree,node=26,state="3")
> # now let's plot using plotSimmap to ensure
> # that the correct branches were painted
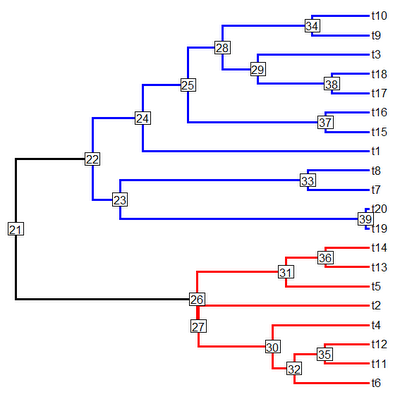
> cols<-c("black","blue","red"); names(cols)<-1:3
> plotSimmap(tree,cols,pts=F,lwd=3,node.numbers=T)
Finally, let's simulate some data (normally, of course, we would read our data from file) and plot a traitgram using the phenogram function in phytools:
> x<-fastBM(tree)
> phenogram(tree,x,colors=cols,fsize=0.8)
That's it.

This comment has been removed by the author.
ReplyDeleteIs there any way to adjust the order the colors plot in so a certain group plots below another? I am working on some data for rodents where roughly half of my taxa are myomorphs of some kind (N = 250) but I am interested in the behavior of the trait for other groups of rodents like squirrels and guinea pigs, which are much less numerous. However, what happens when I plot the tree is the branches for myomorphs are so dense that they completely blot out the less speciose branches of the tree. There is clearly an order to how the groups are plotted because some groups are on top of the others, but re-arranging the order of the labelling factors doesn't appear to change anything. I was wondering if there was any way to plot my most speciose group in the back and plot the other groups on top of them.
ReplyDeletegreat solution!. I am wondering how can I easily extend the paint of the branches to the tips (species names) in the phenogram?
ReplyDelete