A user recently made the following request:
I was just wondering if you would mind posting a short example of the code and data format used to specify colours in the phylomorphospace function (e.g., to create the figure form the paper describing the packages)?
Ok, just for reference, a screen grab of that figure is here:
What this figure shows is a projection of the phylogeny of Caribbean Anolis ecomorph species into a two dimensional morphospace defined by PC 1 & PC 2 from the morphological dataset and tree of Mahler et al. (2010). The tip nodes are colored by ecomorph category; and the internal nodes are black.
All right, for simplicity I'll assume that we want to replicate this general plot in a hypothetical case. First, let's simulate a tree and data for two characters:
> require(phytools)
> tree<-pbtree(n=21)
> X<-cbind(fastBM(tree),fastBM(tree))
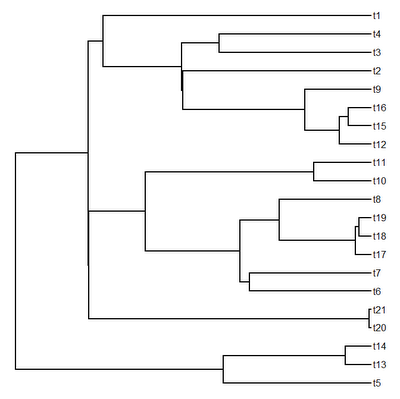
Here, I have just simulated the tree & data; normally we would probably read these from file using functions like read.tree or read.nexus and read.csv. Here is a visualization of our simulated tree:
> plotTree(tree,pts=F)
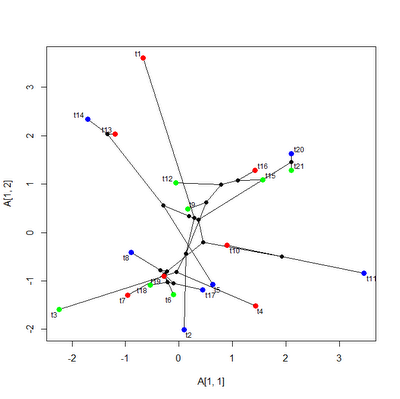
Next, let's say we want to color (for simplicity) every third tip red, blue, or green. (I.e., the series t1, t4, t7, etc. is red.) Normally, we would probably read this from a file as well. For instance, in the ecomorph example, above, we would read a data file containing the ecomorph identification of each of the species from the tree. We could then settle on a color scheme for translation and convert our ecomorph codes to colors. In our example here, we can just do:
> tip.cols<-rep(c("red","blue","green"),7)
> names(tip.cols)<-tree$tip.label
> cols<-c(tip.cols[tree$tip.label],rep("black",tree$Nnode))
> names(cols)<-1:(length(tree$tip)+tree$Nnode)
> phylomorphospace(tree,X,control=list(col.node=cols))
If you examine this figure closely, you'll see that (as intended) everything third numbered tip node is colored red, blue, and green. Obviously, with a real (rather than simulated) tree and dataset we would read in the states for the tip nodes from a file (and they would most likely mean something). It is important to both reorder the color vector by tree$tip.label and re-number it by node number. Don't forget to add elements for internal nodes, even if they are to be colored black (for now at least: perhaps I will fix this in a future version). If we just want to color tip nodes, this is not difficult because the node numbers of the tips are 1 through N (for N species) in the order given by tree$tip.label. It is also probably a good idea to provide X as a matrix & control$col.node as a named vector, just to be safe.
I hope this helps!


Hi Liam!
ReplyDeleteIs there a way to use getDescendents() to color the tips that descend from a certain node (including or excluding the node colors) to use in phylomorphospace()? And if there are several clades one would like to color, how to use the objects of getDescendents for the different clades in the same phylomorphospace?
Thanks for any advice!
Ricardo
I did try reading in the list as you suggested above but had no success.
ReplyDeleteAnswered here. I hope this helps. Liam
DeleteWorks perfect! I tried the phenogram with tidy labels and also works perfect.
ReplyDeleteThanks a lot!
Terrific. Glad to hear it.
DeleteHi Liam,
ReplyDeletehas this function been somehow changed to any extent. I've just upgraded to the most recent version of phytools, and now when I attempt to specify node colors when plotting, I get no colored nodes on the plot (using the exact same script and data that worked previously)? No error messages or anything informative appear when this happens. Any insights appreciated
Can you post which version of phytools you are using (check with packageVersion("phytools")). There is a known bug in the latest CRAN release (described here), but I don't know of any problems (yet) in the most recent non-CRAN version (phytools 0.2-64). Thanks!
DeleteHi Liam,
DeleteI'm using phytools version 0.2-50 (for Windows). I did notice when I load the phytools library file I get the following message:
'Loading required package: rgl
Warning messages:
1: replacing previous import ‘as.igraph’ when loading ‘igraph’
2: replacing previous import ‘edges’ when loading ‘igraph’ '
Any suggestions appreciated
Hi. This is just because phytools dependency 'ape' and a dependency of 'phangorn' ('igraph') have conflicting function names. Basically unless you intend to use those function (as.igraph or edges) - don't worry about it. - Liam
DeleteAlso - there is a newer CRAN version of phytools.
DeleteHi Liam,
ReplyDeleteI can't seem to find a newer Windows version of phytools than 0.2-50. I'm sure this is the same issue someone identified a few weeks ago (somehow I missed it in the blog).
I understand that the Windows binary has been built & this will be available through CRAN within 25 hrs. or so. Thanks! Liam
DeleteHi Liam,
ReplyDeletethat did the trick, thanks for the advice.
cheers
Is there a way to use open circles for the tips?
ReplyDeleteAmazing, thank you!!
ReplyDelete