An R-sig-phylo list serve subscriber asked the following:
I was wondering if there is a method to test for varying rates over time.... I was thinking... in terms of testing if there is a different rate for the entire tree after a specified point in time. For instance, if a snail predator colonizes an island 3.4 Mya, is there evidence for an increased rate of evolution in the prey after that point in time?
Matt Pennell, a Ph.D student in the Harmon lab, responded that this could easily be done using the functions of phytools. He's right, and here's a quick primer (surprisingly pronounced "primmer", for some reason, in American English).
> library(phytools)
> # now let's simulate a stochastic tree
> tree<-pbtree(n=100,scale=15)
> # let's map two states on the tree
> # one for the time before 3.4mybp
> tree<-make.era.map(tree,c(0,max(nodeHeights(tree))-3.4))
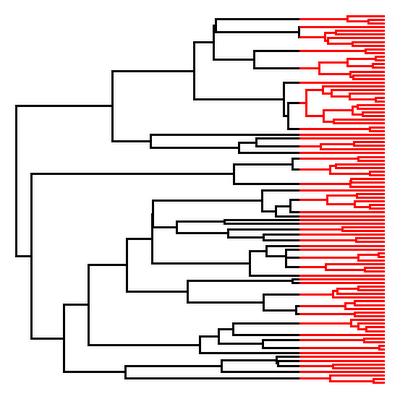
> plotSimmap(tree,pts=F,lwd=3,ftype="off") # visualize
> # now let's simulate some data (since we don't have any)
> x<-sim.rates(tree,c(1,10))
names absent from sig2: assuming same order as $mapped.edge
> # finally, let's fit our two rate model
> fit<-brownie.lite(tree,x)
> fit
$sig2.single
[1] 8.750253
...
$logL1
[1] -316.6358
...
$sig2.multiple
1 2
0.6825954 11.3658746
...
$logL.multiple
[1] -310.6544
...
$P.chisq
[1] 0.0005427315
...
That's it.
Another great data for my thesis....
ReplyDeleteHi Liam, Matt
ReplyDeleteIs this also doable with a discrete trait? i.e. following the example above, is there evidence for an increased rate in shifts to, say, dry/wet habitats following island colonization 3.4Mya?
Thanks,
Vera
Hi Liam,
ReplyDeleteAs requested by Vera, if we can map changes at a fixed point for several selective regimes, this would be awesome!
Julien