As described in a couple of earlier posts (1, 2) I have been doing some work on the function pbtree which does pure-birth (i.e., Yule process) phylogeny simulations.
Well, as the title of this post would suggest, the function can now do birth-death simulation, as well as pure-birth. Remember, pbtree can now condition on total time, the number of tips, or both (by rejection sampling); and can simulate in continuous or discrete time.
Simulating with death (i.e., extinction) is not too hard. For continuous time simulation, whereas before we drew our waiting times from the exponential distribution with λ = b × m for birth-rate b & number of 'open' lineages m, with extinction we draw our wait times from the exponential distribution with λ = (b + d) × m. This is because if the wait time to an event of class A is exponentially distributed with rate λA & the wait time to an event of class B is exponentially distributed with rate λB, and A & B are independent, then the wait times to A or B are exponentially distributed with λ = λA + λB. The probabilities that an event is a birth or a death given that it has occurred are just b/(b+d) and d/(b+d), respectively. This means that our procedure for growing the tree should be (1) draw a random wait time; (2) pick a lineage at random; and (3) and then have it speciate or die with the probabilities given above.
For discrete time the procedure is only slightly different. Now we draw a random wait time from the geometric distribution with p = b + d.** (**Remember that we are actually drawing m independent waiting times & then using the one or more smallest times.) However, since p is a probability (whereas λ was a rate), we are now constrained such that b + d ≤ 1.0. This is because a given lineage can speciate, go extinct, or do nothing within a time-step - but it can't do more than one of these. (b + d = 1.0 guarantees that an event of some kind - speciation or extinction - occurs in every lineage in every generation.)
Code for the new version of pbtree is here. There is also a new phytools build (phytools 0.2-44) with these updates, and it can be downloaded & installed from source.
Here's a little demo of the function, using continuous time simulations:
Loading required package: phytools
> packageVersion("phytools")
[1] ‘0.2.44’
> # continuous time, taxa stop
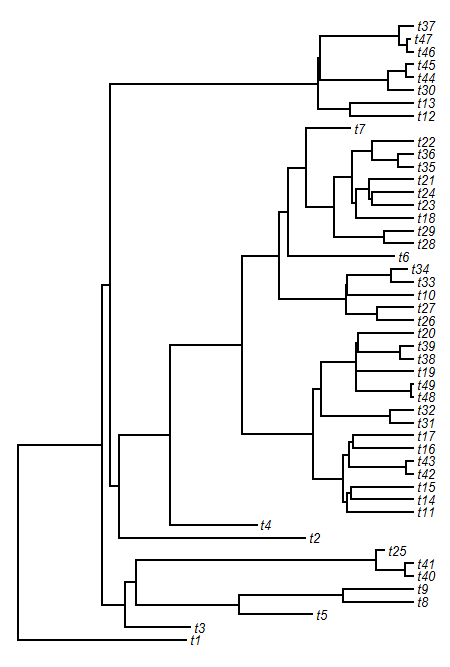
> tree<-pbtree(d=0.2,n=50)
> plotTree(tree,fsize=0.8,ftype="i")
> tree<-pbtree(d=0.2,n=50)
> plotTree(tree,fsize=0.8,ftype="i")
> tt<-log(50)-log(2); tt
[1] 3.218876
> tree<-pbtree(b=1.2,d=0.2,t=tt)
> plotTree(tree,fsize=0.8,ftype="i")
> max(nodeHeights(tree))
[1] 3.218876
> tree<-pbtree(d=0.5,n=100,extant.only=TRUE)
> plotTree(tree,ftype="off")
> ltt(tree,plot=FALSE)$gamma # pull of the recent
[1] 2.298986
> tt # time for simulation
[1] 3.218876
> tree<-pbtree(b=1.2,d=0.2,n=50,t=tt)
simulating with both taxa-stop (n) and time-stop (t) is
performed via rejection sampling & may be slow
13 trees rejected before finding a tree
> max(nodeHeights(tree))
[1] 3.218876
> length(getExtant(tree))
[1] 50
> plotTree(tree,fsize=0.8,ftype="i")
> # function to get the number of extant tips
> ff<-function(x){
tol<-1e-12
if(max(nodeHeights(x))<(tt-tol)) 0
else length(getExtant(x))
}
> # low extinction rate
> lowD<-pbtree(b=1.1,d=0.1,t=tt,nsim=1000)
> lowD
1000 phylogenetic trees
> NlD<-sapply(lowD,ff)
> mean(NlD)
[1] 48.283
> var(NlD)
[1] 1360.986
> # high extinction rate
> highD<-pbtree(b=1.5,d=0.5,t=tt,nsim=1000)
> highD
1000 phylogenetic trees
> NhD<-sapply(highD,ff)
> mean(NhD)
[1] 48.681
> var(NhD)
[1] 2321.631
> # distribution of tree sizes
> hist(NlD,0:ceiling(max(c(NlD,NhD))/10)*10,col="grey", xlab="frequency",ylab="number of tips",main=NULL)
Pretty cool. The purpose of the final exercise was just to illustrate that although the expected number of lineages at time t can be computed as N = e(b-d)×t, the variance among simulations goes up with increasing b.
That's it. I've been testing out the new pbtree extensively - but please report if you are able to break it! Thanks!





No comments:
Post a Comment
Note: due to the very large amount of spam, all comments are now automatically submitted for moderation.